Anti-UBE1L: Mouse UBE1L Antibody |
 |
BACKGROUND Protein degradation is regulated by the ubiquitin-proteasome pathway. Ubiquitin, a 76–amino acid polypeptide, is conjugated to target proteins through isopeptide bonds, resulting in monoubiquitinated and polyubiquitinated substrates. Polyubiquitination marks the substrate protein for proteolysis by the proteasome. Protein ubiquitination is tightly regulated by a ubiquitin-conjugating multienzyme cascade, including the ubiquitin-activating enzyme (E1), ubiquitin conjugases (E2), ubiquitin ligases (E3), and deubiquitinating (DUB) enzymes, which are deconjugases that remove ubiquitin from substrate proteins. UbE1L was identified as the E1-activating enzyme for ISG15. UbE1L is a monomeric protein of 120 kDa. Its most distinctive property is its selectivity in transferring its ISG15 substrate to a single E2 enzyme, UbcH8, an E2 that has been reported to also function in Ub conjugation. The amino acid sequence of UbE1L has a 45% identity with the human ubiquitin-activating enzyme UBE1. The ATP binding site and the active site Cys residue of UBE1 are conserved in UBE1L. The IFN-stimulated gene, 15-kDa protein (ISG15), is a ubiquitin-like protein. Ubiquitin-like proteins conjugate intracellular proteins via an isopeptide linkage. ISG15ylation is reversible and regulated by a similar set of enzymes as in the ubiquitin system. The ubiquitin-activating enzyme-E1-like protein (UBE1L; E1), ubiquitin-conjugating enzyme 8 (E2), estrogen-responsive finger protein (E3), and ubiquitin-specific protease 18 (UBP43 or USP18) have each been reported as enzymes involved in ISG15ylation. At least 200 putative ISG15 target proteins have been identified so far. Many of them have crucial functions in the type I IFN response, and include JAK1, STAT1, RIG-I and the antiviral effector proteins MxA, PKR and RNase L. ISG15 has been reported to prevent virus-mediated degradation of interferon regulatory factor 3 (IRF3), thereby increasing the induction of IFNβ expression. Other reports support a role for ISG15 in mediating resistance to the Ebola virus through ISGylation of NEDD4. Furthermore, ISGylation of target proteins affects their cellular function in the control of IFN-mediated signaling pathway. It was demonstrated that ISGylation of filamin B acts as a negative feedback regulatory gate for the desensitization of type I IFN-induced JNK signaling.1 Moreover, expression of the ISGylation system suppresses NF-k B activation via TRAF6/TAK1 and reduces the level of polyubiquitinated TRAF6. Thus, ISGylation may negatively control the NF-k B pathway.2 In addition, it was shown that UBE1L-ISG15 acts as a distinct growth-suppressive pathway exerting antineoplastic effects by targeting cyclin D1 for repression.3
REFERENCES
1. Joen, Y.J. et al: EMBO Report 10:374-80, 2009
2. Minakawa, M. et al: Biol. Pharm. Bull. 31:2223-7, 2008
3. Feng, Q. et al: Mol. Cancer Therap. 7:3780-8, 2008
2. Minakawa, M. et al: Biol. Pharm. Bull. 31:2223-7, 2008
3. Feng, Q. et al: Mol. Cancer Therap. 7:3780-8, 2008
Products are for research use only. They are not intended for human, animal, or diagnostic applications.
Параметры
Cat.No.: | CP10259 |
Antigen: | Purified recombinant human UBE1L fragments expressed in E. coli. |
Isotype: | Mouse IgG1 |
Species & predicted species cross- reactivity ( ): | Human, Mouse, Rat |
Applications & Suggested starting dilutions:* | WB 1:1000 IP 1:50 IHC n/d ICC n/d FACS n/d |
Predicted Molecular Weight of protein: | 120 kDa |
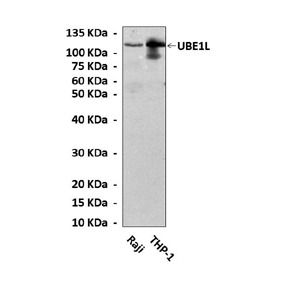
Specificity/Sensitivity: | Detects endogenous UBE1L proteins without cross-reactivity with other related proteins. |
Storage: | Store at -20°C, 4°C for frequent use. Avoid repeated freeze-thaw cycles. |
*Optimal working dilutions must be determined by end user.
Документы
Информация представлена исключительно в ознакомительных целях и ни при каких условиях не является публичной офертой