Anti-Ask1/MKK5: Mouse Ask1/MKK5 Antibody |
 |
BACKGROUND MAPK cascades, which include ERK1/2, JNK, the big MAPK/ERK5, and the p38 MAPK cascades, are intracellular signaling networks utilized by mammalian cells to respond to a wide spectrum of extracellular stimuli. MAPKs are activated through a module that includes an MAPK, an MAPK kinase (MAPKK), and an MAPKK kinase (MAP3K). Located upstream in the MAPK module, an MAP3K phosphorylates and activates its downstream MAPKKs, which in turn phosphorylate and activate the downstream MAPKs. A growing number of MAP3Ks including MEKK1, MEKK2, MEKK3, MEKK4, apoptosis signal-regulating kinase 1 ASK1, transforming growth factor-beta-activated kinase 1 (TAK1), dual leucine-zipper kinase, and tumor progression locus 2 (Tpl2) have been identified that are capable of activating multiple MAPKs. Because of its unique position in the MAPK module, a MAP3K is crucial in relaying the upstream receptor-mediated signals through the MAPK cascades to induce physiological responses. Biochemical and genetic studies have demonstrated that these MAP3Ks are crucial in relaying distinct cell-surface signals through various downstream MAPK pathways to their cytoplasmic and nuclear effectors. Studies using gene-targeting mice have also revealed that MAP3Ks have both overlapping and distinctive functions. For instance, many MAP3Ks also activate the IkappaB kinase (IKK)–nuclear factor kappaB (NF-kappaB) pathway.1
Apoptosis signal-regulating kinase 1 (ASK1)/MEKK5 is one of several MAP3Ks that are activated in response to proinflammatory stimuli, hydrogen peroxide (H2O2) and cellular stress, leading to activation of both the JNK/SAPKs (via activation of SEK1) and the p38s (via activation of MKK3 and MKK6). ASK1 is a critical mediator in TNF, ROS and stress-induced cell death. It was shown that ASK1 could physically associate with adapter polypeptides that transduce signals from TNF receptor 1 (TNFR1). Thus, the carboxyl-terminal noncatalytic domain of ASK1 has been shown to interact in vivo with TNFR-associated factor 2 (TRAF2), an adapter protein that is required for coupling TNFR1 to the SAPKs. ASK1 can also associate with TRAF5 and -6, additional TRAFs implicated in the regulation of the SAPKs by members of the TNF superfamily. Activation of ASK1 by TNF requires the ROS-mediated dissociation of Trx possibly followed by the binding of TRAF2 and consequent ASK1 homo-oligomerization.2 ASK1 has also been shown to bind and be activated by Daxx, an adapter protein originally thought to relay signals from Fas to the SAPKs. However, establishment of a function for Daxx has been controversial. ASK1 is reported to induce apoptosis by triggering a mitochondria-dependent pathway including Bid cleavage, Bax mitochondria translocation, cytochrome c release from mitochondria and subsequent activation of caspase 9 and caspase 3.3
ASK1 is a central target of many cellular survival factors that bind to different domains of ASK1 to keep ASK1 in an inactive state. Redox sensor proteins thioredoxin (Trx) binds to the N-terminal domain, while glutaredoxin binds to the C-terminal domain of ASK1 to inhibit ASK1 kinase activity. Phosphoserine-binding protein 14-3-3 associates with ASK1 via the pSer967 site of ASK1 and inhibits ASK1-induced apoptosis. Several proteins such as heat–shock protein hsp72 block ASK1 oligomerization and recruitment of its downstream targets. ASK1 can be phosphorylated at several sites and these phosphorylation sites regulate ASK1 activity in both positive and negative manner. Thus, protein phosphatase 5 (PP5) dephosphorylates ASK1 at the autophosphorylation site pThr845 to function as a negative feedback inhibitor of ASK1 signaling. Conversely, several serine/threonine protein kinases such as Akt directly phosphorylates ASK1 at Ser83 to inhibit ASK1-induced apoptosis.4
REFERENCES
1. Seger,R. & Krebs,E.G.: FASEB J. 9:726-35, 1995
2. Liu, H. et al: Mol. Cell. Biol. 20:2198-208, 2000
3. Matsukawa, J. et al: J. Biochem. 136:261-5, 2004
4. Yuan, Z-Q. et al: J. Biol. Chem. 278:23432-40, 2003
2. Liu, H. et al: Mol. Cell. Biol. 20:2198-208, 2000
3. Matsukawa, J. et al: J. Biochem. 136:261-5, 2004
4. Yuan, Z-Q. et al: J. Biol. Chem. 278:23432-40, 2003
Products are for research use only. They are not intended for human, animal, or diagnostic applications.
Параметры
Cat.No.: | CP10395 |
Antigen: | Recombinant human Ask1 fragments expressed in E. coli. |
Isotype: | Mouse IgG1 |
Species & predicted species cross- reactivity ( ): | Human |
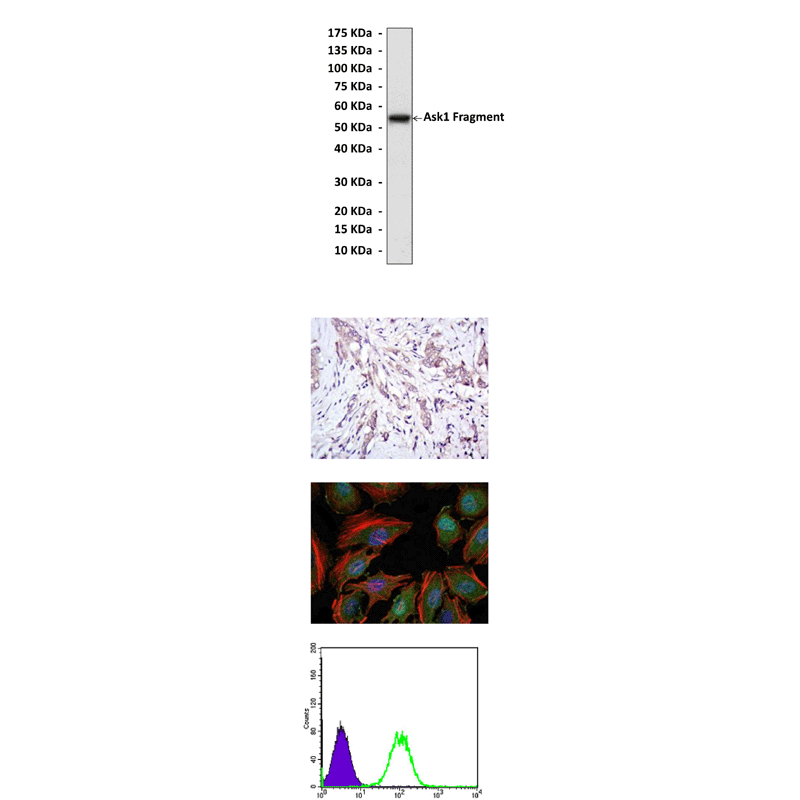
Applications & Suggested starting dilutions:* | WB 1:1000 IP n/d IHC 1:50 - 1:200 ICC 1:50 - 1:200 FACS 1:50 - 1:200 |
Predicted Molecular Weight of protein: | 170 kDa |
Specificity/Sensitivity: | Detects Ask1 proteins in various cell lysate. |
Storage: | Store at -20°C, 4°C for frequent use. Avoid repeated freeze-thaw cycles. |
*Optimal working dilutions must be determined by end user.
Информация представлена исключительно в ознакомительных целях и ни при каких условиях не является публичной офертой