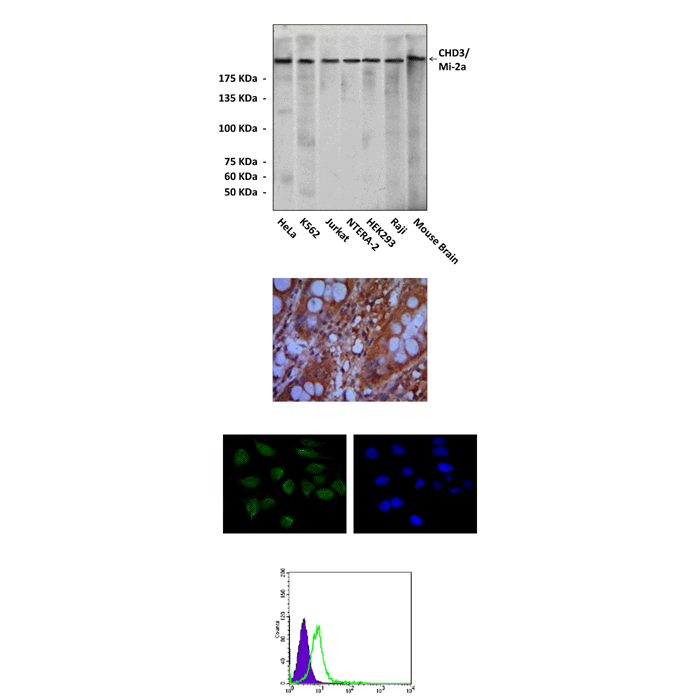
Anti-CHD3/Mi-2a: Mouse CHD3/Mi-2a Antibody |
 |
BACKGROUND Dynamic regulation of chromatin structure is an important mechanism for modulating genome activity in eukaryotes. Two major groups of proteins with enzymatic activities directed toward chromatin are histone-modifying enzymes and ATP-dependent chromatin remodelers. The former modify histones by attaching or removing acetyl, methyl, phosphate, ADP-ribose, ubiquitin, or SUMO groups. These modifications alone do not lead to major changes of chromatin structure. The histone code hypothesis postulates that these "marks" are specifically bound by factors which in turn influence chromatin structure by mechanisms that are not well understood. ATP-dependent chromatin remodelers can directly change nucleosome structure by altering interactions between DNA and the histone octamer. This results in nucleosome assembly or eviction, incorporation of histone variants, altered DNA-to-octamer contacts within an existing nucleosome, or lateral movement of the octamer along DNA. As a result, the accessibility of nucleosomal DNA is changed. Most ATP-dependent chromatin remodelers are multisubunit complexes containing an ATPase of the SNF2 superfamily. These enzymes share a conserved ATPase domain but can differ in additional domains. Accordingly, they are subdivided into several families, including the DDM1, the SWI/SNF, the ISWI, the INO80, and the CHD family.1
CHD proteins are characterized by the presence of two tandemly arranged chromodomains. The mammalian CHD family has nine members. The CHD3(Mi-2a)/CHD4 (Mi-2b) subfamily is characterized by the presence of a pair of PHD fingers preceding the chromodomains. These proteins exist in multisubunit complexes with nucleosome remodeling and deacetylation (NuRD) activity. In addition to Chd3 or Chd4, NuRD complexes contain histone deacetylase 1 (HDAC1) and HDAC2, the histone binding proteins RbAp46 and RbAp48, proteins of the MTA and p66 families (MTA1, MTA2, or MTA3 and p66 or p66β), and subunits with methylated DNA binding domains (MBD3). As part of complex, CHD3 directly interacts with HDAC1 and 2, ATR, and TRIM27. It has been suggested that these complexes couple ATP-dependent chromatin remodeling and deacetylation to effect transcriptional repression. However, several studies also implicate a function of CHD3 family members in transcriptional activation.2 Complexes related to NuRD exist in Drosophila melanogaster and Caenorhabditis elegans and play important roles during differentiation and cell fate determination. This function is conserved in mammals: an MTA3-containing NuRD complex regulates B-lymphocyte differentiation and is indispensable for BCL-6-mediated repression of plasma cell-specific genes in B lymphocytes. Two different modes of targeting NuRD complexes have been suggested. First, recruitment via interaction with DNA bound transcriptional repressors, such as BCL-6, Hunchback, or Tramtrack 69. Second, direct binding of stoichiometric (MBD3) or associated (MBD2A) MBD proteins to methylated DNA.3 Furthmore, CK2 phosphorylation is a common feature of CHD family ATPases.4 CHD proteins play important role in the regulation of gene expression, recombinantion, genome organization, and chromatin structure, as well as their potential activity in RNA processing. In addition, CHD proteins are also implicated in human disease.
CHD proteins are characterized by the presence of two tandemly arranged chromodomains. The mammalian CHD family has nine members. The CHD3(Mi-2a)/CHD4 (Mi-2b) subfamily is characterized by the presence of a pair of PHD fingers preceding the chromodomains. These proteins exist in multisubunit complexes with nucleosome remodeling and deacetylation (NuRD) activity. In addition to Chd3 or Chd4, NuRD complexes contain histone deacetylase 1 (HDAC1) and HDAC2, the histone binding proteins RbAp46 and RbAp48, proteins of the MTA and p66 families (MTA1, MTA2, or MTA3 and p66 or p66β), and subunits with methylated DNA binding domains (MBD3). As part of complex, CHD3 directly interacts with HDAC1 and 2, ATR, and TRIM27. It has been suggested that these complexes couple ATP-dependent chromatin remodeling and deacetylation to effect transcriptional repression. However, several studies also implicate a function of CHD3 family members in transcriptional activation.2 Complexes related to NuRD exist in Drosophila melanogaster and Caenorhabditis elegans and play important roles during differentiation and cell fate determination. This function is conserved in mammals: an MTA3-containing NuRD complex regulates B-lymphocyte differentiation and is indispensable for BCL-6-mediated repression of plasma cell-specific genes in B lymphocytes. Two different modes of targeting NuRD complexes have been suggested. First, recruitment via interaction with DNA bound transcriptional repressors, such as BCL-6, Hunchback, or Tramtrack 69. Second, direct binding of stoichiometric (MBD3) or associated (MBD2A) MBD proteins to methylated DNA.3 Furthmore, CK2 phosphorylation is a common feature of CHD family ATPases.4 CHD proteins play important role in the regulation of gene expression, recombinantion, genome organization, and chromatin structure, as well as their potential activity in RNA processing. In addition, CHD proteins are also implicated in human disease.
REFERENCES
1. Hall, J.A. & Georgel, P.T.: Biochem. Cell. Biol. 85:463-76, 2007
2. Marfella, C.G.A. & Imbalzano, A.N.: Mutat. Res./Fundament. Mol. Mech. Mutagen.618:30-40, 2007
3. Vignali, M. et al: Mol. Cell. Biol. 20:1899-1910, 2000
4. Bouazoune, K. & Brehm, A.: J. Biol. Chem. 280:41912-20, 2005
2. Marfella, C.G.A. & Imbalzano, A.N.: Mutat. Res./Fundament. Mol. Mech. Mutagen.618:30-40, 2007
3. Vignali, M. et al: Mol. Cell. Biol. 20:1899-1910, 2000
4. Bouazoune, K. & Brehm, A.: J. Biol. Chem. 280:41912-20, 2005
Products are for research use only. They are not intended for human, animal, or diagnostic applications.
Параметры
Cat.No.: | CP10302 |
Antigen: | Raised against recombinant human CHD3 fragments expressed in E. coli. |
Isotype: | Mouse IgG1 |
Species & predicted species cross- reactivity ( ): | Human, Mouse, Rat |
Applications & Suggested starting dilutions:* | WB 1:1000 IP n/d IHC 1:50 - 1:200 ICC 1:50 - 1:200 FACS 1:50 - 1:200 |
Predicted Molecular Weight of protein: | 226 kDa |
Specificity/Sensitivity: | Detects endogenous CHD3 proteins without cross-reactivity with other family members. |
Storage: | Store at -20°C, 4°C for frequent use. Avoid repeated freeze-thaw cycles. |
*Optimal working dilutions must be determined by end user.
Документы
Информация представлена исключительно в ознакомительных целях и ни при каких условиях не является публичной офертой