Anti-Phospho-IκB-α: Rabbit IkappaB-alpha, Phospho-Ser32 Antibody |
 |
BACKGROUND The IkappaBs form a small family of related proteins with a core consisting of six or more ankyrin repeats, an N-terminal regulatory domain and a C-terminal domain that contains a PEST motif. The best-studied and major IkappaB protein is IkappaBalpha and IkappaBbeta. Members of the IkappaB protein family are constitutively and ubiquitously expressed proteins that localize in the cytosol, except for Bcl-3 and IkappaBzeta which are primarily present in the nucleus. The main function of IkappaB proteins is to inhibit NF-kappaB activity when it is not required, and this happens via protein-protein interaction that takes place between IkappaB proteins and NF-kappaB dimers in the cytosol. IkappaBalpha and IkappaBbeta interact via a tertiary structure formed by their ANK repeats and C-terminal region (i.e. PEST domain) with the RHD of NF-kappaB dimerized proteins in such a way that masks the positively charged regions of the NLSs within the RHDs of NF-kappaB dimers. As a result, NF-kappaB dimers are prevented from translocating to the nucleus, and thus, they are kept in an inactive, IkappaB-bound state in the cytosol.1 In addition, It is known that NF-kappaB is itself an upregulator of IkappaBalpha and IkappaBbeta. Induced expression of IkappaBalpha allows translocation of nascently synthesized IkappaBalpha into the nucleus, where it binds to active NF-kappaB dimers that are bound to kappaB sites within the promoters of their target genes, and leads to dissociation between NF-kappaB dimers and DNA. It forces a conformational change in IkappaBalpha that exposes the nuclear export signal (NES), eventually leading to resequesterization of IkappaBalpha and NF-kappaB dimers in the cytosol.2
Activation of NF-kappaB correlates with phosphorylation of IkappaBalpha and requires the proteolysis of this inhibitor. The phosphorylation of IkappaBs was catalyzed by IkappaB kinases (IKKs). IKK complex with catalytically active IKKalpha and IKKbeta is responsible for phosphorylating two N-terminally located conserved serine residues in IkappaBalpha and IkappaBbeta (Ser32 and Ser36 in IkappaBalpha; Ser19 and Ser23 in IkappaBbeta. Phosphorylation of these sites renders them susceptible to ubiquitination and subsequent proteolytic degradation. After signal-induced phosphorylation of IkappaBalpha on serine residues 32 and 36, lysine residues 21 and 22 are major sites of ubiquitin ligation which target the protein for rapid degradation by the proteasome.3 In addition, the sumoylated form of IkappaBalpha was found to be highly resistant to signal-induced ubiquitination. Indeed, it was demonstrated that SUMO-4-mediated downregulation of NF-kappaB was shown to be dependent on modification of IkappaBalpha by SUMO-4.4
In sum, for signal-induced NF-kappaB transactivation activity to manifest, at least six main biochemical events must precede: (1) phosphorylation of IkappaBalpha and IkappaBbeta by the IKK complex, (2) ubiquitination of phosphorylated IkappaBalpha, (3) proteasome-mediated degradation of IkappaBalpha and IkappaBbeta, (4) phosphorylation of NF-kappaB dimer, (5) nuclear translocation of NF-kappaB dimer, and (6) NF-kappaB dimer-kappaB DNA interaction.
Activation of NF-kappaB correlates with phosphorylation of IkappaBalpha and requires the proteolysis of this inhibitor. The phosphorylation of IkappaBs was catalyzed by IkappaB kinases (IKKs). IKK complex with catalytically active IKKalpha and IKKbeta is responsible for phosphorylating two N-terminally located conserved serine residues in IkappaBalpha and IkappaBbeta (Ser32 and Ser36 in IkappaBalpha; Ser19 and Ser23 in IkappaBbeta. Phosphorylation of these sites renders them susceptible to ubiquitination and subsequent proteolytic degradation. After signal-induced phosphorylation of IkappaBalpha on serine residues 32 and 36, lysine residues 21 and 22 are major sites of ubiquitin ligation which target the protein for rapid degradation by the proteasome.3 In addition, the sumoylated form of IkappaBalpha was found to be highly resistant to signal-induced ubiquitination. Indeed, it was demonstrated that SUMO-4-mediated downregulation of NF-kappaB was shown to be dependent on modification of IkappaBalpha by SUMO-4.4
In sum, for signal-induced NF-kappaB transactivation activity to manifest, at least six main biochemical events must precede: (1) phosphorylation of IkappaBalpha and IkappaBbeta by the IKK complex, (2) ubiquitination of phosphorylated IkappaBalpha, (3) proteasome-mediated degradation of IkappaBalpha and IkappaBbeta, (4) phosphorylation of NF-kappaB dimer, (5) nuclear translocation of NF-kappaB dimer, and (6) NF-kappaB dimer-kappaB DNA interaction.
REFERENCES
1. Karin, M.: Oncogene 18:6867-74, 1999
2. Arenzana-Seisdedos, F. et al: Mol. Cell. Biol. 15:2689-96, 1995
3. Rodriguez, M.S. et al: Oncogene 12:2425-35, 1996
4. Guo, D. et al: Nature Genet.26:837-41, 2004
2. Arenzana-Seisdedos, F. et al: Mol. Cell. Biol. 15:2689-96, 1995
3. Rodriguez, M.S. et al: Oncogene 12:2425-35, 1996
4. Guo, D. et al: Nature Genet.26:837-41, 2004
Products are for research use only. They are not intended for human, animal, or diagnostic applications.
Параметры
Cat.No.: | CG1242 |
Antigen: | Short peptide from human IKappaB-alpha sequence surrounding and containing phospho-Ser32. |
Isotype: | Rabbit IgG |
Species & predicted species cross- reactivity ( ): | Human, Mouse, Rat |
Applications & Suggested starting dilutions:* | WB 1:1000 - 1:10000 IP 1:50 - 1:100 IHC n/d ICC 1:50 - 1:100 FACS n/d |
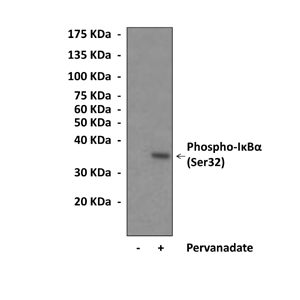
Predicted Molecular Weight of protein: | 35 kDa |
Specificity/Sensitivity: | Detects endogenous phosphorylated IKappaB-alpha (Ser32) proteins without cross-reactivity with other family members. |
Storage: | Store at -20°C, 4°C for frequent use. Avoid repeated freeze-thaw cycles. |
*Optimal working dilutions must be determined by end user.
Информация представлена исключительно в ознакомительных целях и ни при каких условиях не является публичной офертой