Anti-Rictor: Mouse Rictor Antibody |
 |
BACKGROUND mTOR (mammalian target of rapamycin) is a Ser/Thr kinase that belongs to the phosphatidylinositol (PI) kinase-related protein kinase family. mTOR partitions between two scaffold proteins, Raptor and Rictor. The rapamycin-sensitive Raptor/mTOR complex (TORC1) consists of mTOR, raptor, GβL, and PRAS40, and it functions to regulate protein synthesis and cell growth in response to nutrient levels and growth factor signals via S6K1 and 4EBP1/PHAS. TORC1 is regulated by TSC-Rheb (tuberous sclerosis complex-Ras homolog-enriched in brain) signaling. AMPK acts as an intracellular energy sensor that controls glucose and lipid metabolism in peripheral tissues. AMPK- and PI3-kinase/Akt-dependent signaling pathways converge at the level of tuberus sclerosis complex 2 (TSC2 or tuberin) which functions as a GTPase-activating protein for the small GTPase Rheb. Akt phosphorylation of the tuberous sclerosis complex (TSC1 and -2) relieves TSC's GTPase activating protein (GAP) function, allowing the accumulation of Rheb GTP, which then binds mTOR, allowing activation of the raptor complex in a GTP-dependent manner. The activated raptor complex increases translation by phosphorylating its targets. Normal cell growth and homeostasis rely on a proper balance between nutrient- and hormone-dependent regulation of the AMPK/mTOR axis, while its deregulation could lead to metabolic and growth-related diseases.1
Rictor (rapamycin-insensitive companion of mTOR) was the first identified subunit of rapamycin-insensitive Rictor/mTOR (TORC2) complex. It is unique to TORC2. Rictor is the homolog of AVO3 in yeast. TORC2 is also called the Rictor complex. Rictor is a large protein with a predicted molecular weight of 190 kDa. It has some domain structures in the amino terminal region that are relatively conserved among species, but the functions of these domains are not known. It is speculated that these domains may mediate substrate binding and are important for TORC2 assembly. The interaction between Rictor and mTOR is not blocked by rapamycin nor affected by nutrient levels, which are conditions known to regulate TORC1. Thus, it is not surprising that knockdown of Rictor by RNAi in cultured cells does not change the phosphorylation status of S6K1 and 4EBP1. This suggests that TORC2 has different physiological functions from TORC1. The overall physiological importance of Rictor is emphasized by the fact that the Rictor knockout mice die around E10.5, possibly due to defects in vascular development
The rapamycin-insensitive Rictor/mTOR (TORC2) complex consists of mTOR, rictor, GβL, Sin1, PRR5, and protor.2,3 TORC2 regulates cell growth and survival in response to hormonal signals. TORC2 is activated by growth factors, but, in contrast to mTORC1, seems to be nutrient-insensitive. TORC2 seems to function upstream of Rho GTPases to regulate the actin cytoskeleton, probably by activating one or more Rho-type guanine nucleotide exchange factors. TORC2 promotes the serum-induced formation of stress-fibers or F-actin. TORC2 plays a critical role in AKT Ser-473 phosphorylation, which may facilitate the phosphorylation of the activation loop of AKT1 on Thr-308 by PDK1 which is a prerequisite for full activation. TORC2 regulates the phosphorylation of SGK1 at Ser-422. TORC2 also modulates the phosphorylation of PRKCA on Ser-657. It plays an essential role in embryonic growth and development. In certain cell types, prolonged inhibition of mTOR by rapamycin may impair mTORC2 assembly and hence AKT activation.4 TORC2 is downstream of PI3K signaling. Moreover, it was reported that Rheb exerts a negative effect on TORC2 probably through the S6K1-dependent negative feedback loop.5
Rictor (rapamycin-insensitive companion of mTOR) was the first identified subunit of rapamycin-insensitive Rictor/mTOR (TORC2) complex. It is unique to TORC2. Rictor is the homolog of AVO3 in yeast. TORC2 is also called the Rictor complex. Rictor is a large protein with a predicted molecular weight of 190 kDa. It has some domain structures in the amino terminal region that are relatively conserved among species, but the functions of these domains are not known. It is speculated that these domains may mediate substrate binding and are important for TORC2 assembly. The interaction between Rictor and mTOR is not blocked by rapamycin nor affected by nutrient levels, which are conditions known to regulate TORC1. Thus, it is not surprising that knockdown of Rictor by RNAi in cultured cells does not change the phosphorylation status of S6K1 and 4EBP1. This suggests that TORC2 has different physiological functions from TORC1. The overall physiological importance of Rictor is emphasized by the fact that the Rictor knockout mice die around E10.5, possibly due to defects in vascular development
The rapamycin-insensitive Rictor/mTOR (TORC2) complex consists of mTOR, rictor, GβL, Sin1, PRR5, and protor.2,3 TORC2 regulates cell growth and survival in response to hormonal signals. TORC2 is activated by growth factors, but, in contrast to mTORC1, seems to be nutrient-insensitive. TORC2 seems to function upstream of Rho GTPases to regulate the actin cytoskeleton, probably by activating one or more Rho-type guanine nucleotide exchange factors. TORC2 promotes the serum-induced formation of stress-fibers or F-actin. TORC2 plays a critical role in AKT Ser-473 phosphorylation, which may facilitate the phosphorylation of the activation loop of AKT1 on Thr-308 by PDK1 which is a prerequisite for full activation. TORC2 regulates the phosphorylation of SGK1 at Ser-422. TORC2 also modulates the phosphorylation of PRKCA on Ser-657. It plays an essential role in embryonic growth and development. In certain cell types, prolonged inhibition of mTOR by rapamycin may impair mTORC2 assembly and hence AKT activation.4 TORC2 is downstream of PI3K signaling. Moreover, it was reported that Rheb exerts a negative effect on TORC2 probably through the S6K1-dependent negative feedback loop.5
REFERENCES
1. Hay, A. & Sonenberg, N.: Gene Dev. 18:1926-45, 2004
2. Woo, S-Y. et al: J. Biol. Chem. 282:25604-12, 2007
3. Pearce, L.R. et al: Biochem J. 405(Pt 3): 513–522, 2007
4. Cybulski, N. & Hall, M.N.:Trends Biochem Sci. 34: 620-627, 2009
5. Yang, Q. et al: Proc. Natl. Acad. Sci. USA 103:6811-6, 2006
2. Woo, S-Y. et al: J. Biol. Chem. 282:25604-12, 2007
3. Pearce, L.R. et al: Biochem J. 405(Pt 3): 513–522, 2007
4. Cybulski, N. & Hall, M.N.:Trends Biochem Sci. 34: 620-627, 2009
5. Yang, Q. et al: Proc. Natl. Acad. Sci. USA 103:6811-6, 2006
Products are for research use only. They are not intended for human, animal, or diagnostic applications.
Параметры
Cat.No.: | CP10256 |
Antigen: | Purified recombinant human Rictor fragments expressed in E. coli. |
Isotype: | Mouse IgG1 |
Species & predicted species cross- reactivity ( ): | Human, Mouse, Rat |
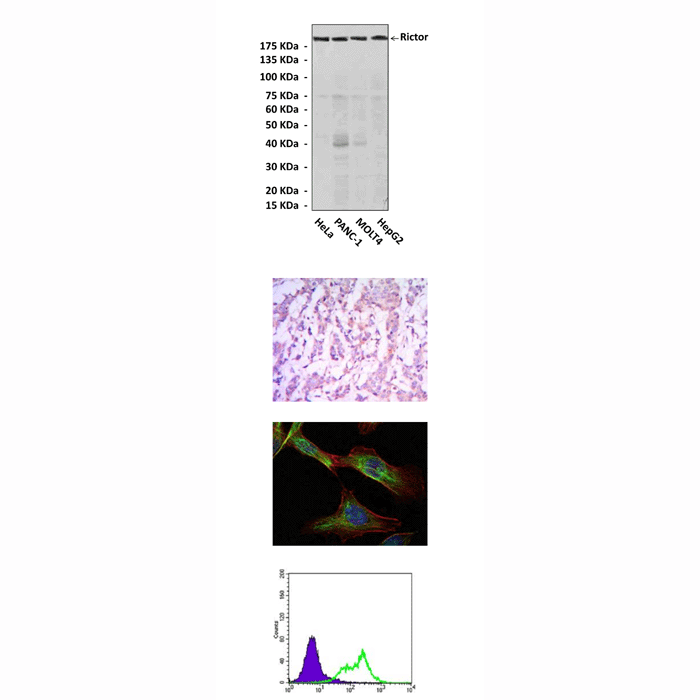
Applications & Suggested starting dilutions:* | WB 1:1000 IP n/d IHC 1:50 - 1:200 ICC 1:50 - 1:200 FACS 1:50 - 1:200 |
Predicted Molecular Weight of protein: | 193 kDa |
Specificity/Sensitivity: | Detects endogenous Rictor proteins without cross-reactivity with other related proteins. |
Storage: | Store at -20°C, 4°C for frequent use. Avoid repeated freeze-thaw cycles. |
*Optimal working dilutions must be determined by end user.
Документы
Информация представлена исключительно в ознакомительных целях и ни при каких условиях не является публичной офертой