Anti-Runx1: Mouse Runx1 Antibody |
 |
BACKGROUND Runx1/AML1 (also known as CBFA2 and PEBP2αB) is a Runt family transcription factor critical for normal hematopoiesis. Runx1 directly binds the enhancer core DNA sequence TGT/cGGT, which is present in a number of different viral and cellular promoters and enhancers. Runx1 binds DNA via a central 118 amino acid Runt homology domain (RHD), and its DNA binding affinity is increased by heterodimerization through the RHD with core-binding factor beta (CBFbeta).1 Runx1 enhances gene transcription by interacting with transcriptional coactivators such as p300 and CREB-binding protein. However, Runx1 can also suppress gene transcription by interacting with transcriptional corepressors, including mSin3A, TLE (mammalian homolog of Groucho), and histone deacetylases. Runx1 not only is critical for definitive hematopoiesis in the fetus but also is required for normal megakaryocytic maturation and T-lymphocyte and B-lymphocyte development in adult mice. Moreover, Runx1 binds the enhancer core motif that is present in the transcriptional regulatory region of a number of different hematopoietic-specific genes, including the T cell antigen receptors, the myeloid-specific gene myeloperoxidase, and a variety of cytokines and their receptors, including granulocyte–macrophage colony-stimulating factor (GM-CSF), interleukin-3 (IL-3), and the colony-stimulating factor 1 (CSF1) receptor. The enhancer core motif is necessary for the tissue-specific expression of these genes, but cannot confer tissue specificity to a heterologous promoter. In addition, this sequence is in the enhancers of the simian virus 40, polyoma virus, and all murine type C retroviruses and has been shown to represent a critical regulatory region in several murine leukemia retroviral enhancers, where it influences tissue tropism and disease phenotype. Thus, Runx1 may function as a transcriptional organizer that recruits tissue-specific factors to stimulate lineage-restricted transcription.Runx1 is likely to play a pivotal role in the regulation of target genes involved in the function of the hematopoietic lineages.2
Runx1 is the most frequent target of chromosomal translocations in acute leukemias. Several chimeric proteins have been identified, including t(8;21) (Runx1-ETO/MTG8), t(16;21) (Runx1-MTG16), t(3;21) (Runx1 -Evi1), t(12;21) (TEL-Runx1), and t(X;21) (Runx1-Fog2). Some of them have transdominant properties for wild-type Runx1 and acts as transcriptional repressors. The transcriptional repression in Runx1 fusion proteins is mediated by recruitment of nuclear corepressor complex that maintains local histone deacetylation. Inhibition of the expression of Runx1-responsive genes leads to a block in hematopoietic cell differentiation and consequent leukemic transformation. On the other hand, mutations in the Runt domain of the Runx1 are identified in both sporadic acute myeloblastic leukemia (AML) without Runx1 translocation and familial platelet disorder with predisposition to AML. These observations indicate that a decrease in Runx1 dosage resulting from chromosomal translocations or mutations contributes to leukemogenesis. Furthermore, dysregulated chromatin remodeling and transcriptional control appears to be a common pathway in Runx1-associated leukemias that could be an important target for the development of new therapeutic agents. However, expression of the fusion protein is not sufficient by itself to cause leukemia and likely requires additional events for leukemogenesis. Point mutations in a Runx1 allele cause haploinsufficiency and a biallelic null for Runx1, which are associated with familial platelet disorder with a propensity for acute myeloid leukemia (FPD/AML) and AML-M0, respectively. Thus, the correct protein structure and the precise dosage of Runx1 are essential for the maintenance of normal hematopoiesis.3
Runx1 is the most frequent target of chromosomal translocations in acute leukemias. Several chimeric proteins have been identified, including t(8;21) (Runx1-ETO/MTG8), t(16;21) (Runx1-MTG16), t(3;21) (Runx1 -Evi1), t(12;21) (TEL-Runx1), and t(X;21) (Runx1-Fog2). Some of them have transdominant properties for wild-type Runx1 and acts as transcriptional repressors. The transcriptional repression in Runx1 fusion proteins is mediated by recruitment of nuclear corepressor complex that maintains local histone deacetylation. Inhibition of the expression of Runx1-responsive genes leads to a block in hematopoietic cell differentiation and consequent leukemic transformation. On the other hand, mutations in the Runt domain of the Runx1 are identified in both sporadic acute myeloblastic leukemia (AML) without Runx1 translocation and familial platelet disorder with predisposition to AML. These observations indicate that a decrease in Runx1 dosage resulting from chromosomal translocations or mutations contributes to leukemogenesis. Furthermore, dysregulated chromatin remodeling and transcriptional control appears to be a common pathway in Runx1-associated leukemias that could be an important target for the development of new therapeutic agents. However, expression of the fusion protein is not sufficient by itself to cause leukemia and likely requires additional events for leukemogenesis. Point mutations in a Runx1 allele cause haploinsufficiency and a biallelic null for Runx1, which are associated with familial platelet disorder with a propensity for acute myeloid leukemia (FPD/AML) and AML-M0, respectively. Thus, the correct protein structure and the precise dosage of Runx1 are essential for the maintenance of normal hematopoiesis.3
REFERENCES
1. Yamagata, T. et al: Int. J. Hematol. 82:1-8, 2005
2. Okuda, T. et al: Cell 84:321-30, 1996
3. Asou, N.: Crit. Rev. Oncol/Hematol. 45:129-50, 2003
2. Okuda, T. et al: Cell 84:321-30, 1996
3. Asou, N.: Crit. Rev. Oncol/Hematol. 45:129-50, 2003
Products are for research use only. They are not intended for human, animal, or diagnostic applications.
Параметры
Cat.No.: | CP10382 |
Antigen: | Raised against purified recombinant fragments of human Runx1 expressed in E. coli. |
Isotype: | Mouse IgG1 |
Species & predicted species cross- reactivity ( ): | Human |
Applications & Suggested starting dilutions:* | WB 1:1000 IP n/d IHC n/d ICC n/d FACS n/d |
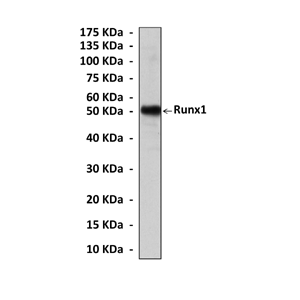
Predicted Molecular Weight of protein: | 50 kDa |
Specificity/Sensitivity: | Detects Runx1 proteins without cross-reactivity with other family members. |
Storage: | Store at -20°C, 4°C for frequent use. Avoid repeated freeze-thaw cycles. |
*Optimal working dilutions must be determined by end user.
Документы
Информация представлена исключительно в ознакомительных целях и ни при каких условиях не является публичной офертой