Anti-PERK: Mouse PERK Antibody |
 |
BACKGROUND Most secreted and transmembrane proteins are synthesized, modified, and folded in the lumen of the endoplasmic reticulum (ER). To ensure the fidelity of protein folding and maturation, cells turn on a network of signaling pathways, collectively termed the unfolded protein response (UPR), to adjust the protein-folding capacity of the ER according to need. Imbalances between protein load and folding capacity is monitored by three distinct UPR sensors: inositol-requiring enzyme 1 (IRE1), protein kinase RNA (PKR)-like ER kinase (PERK), and activating transcription factor-6 (ATF6).1 In mammalian cells, PERK and IRE1 independently govern two key UPR signal transduction pathways. PERK is a transmembrane kinase that phosphorylates translation initiation factor eIF2α, thereby reducing cellular protein synthesis and with it the load of proteins entering into the ER. eIF2α phosphorylation also allows the translation of select mRNAs that contain small open reading frames in their 5′ untranslated regions, leading to the production of transcription activators, such as ATF4 and ATF5. IRE1 is a bifunctional transmembrane serine/threonine kinase/endoribonuclease with an ER–luminal domain that senses misfolded proteins in the ER, most likely by direct ligand-mediated recognition. As a result, IRE1 activates its cytoplasmic kinase and endoribonuclease (RNase) domains, which initiate a nonconventional mRNA splicing reaction that results in the cleavage of the mRNA encoding the transcription factor XBP-1 (x-box binding protein 1), a conserved effector of the UPR that drives the transcription of a plethora of ER-stress responsive genes, including ER resident chaperones and protein modifying enzymes. In addition to splicing Xbp1 mRNA, IRE1's kinase can also activate the c-Jun N-terminal kinase (JNK) signaling pathway through the MAP3K cascade. The transcription factors produced by PERK, IRE1, and other UPR signaling pathways collaborate to control behavior, metabolism, and ultimately cell fate in response to ER stress by inducing a wide array of targets that include protein folding chaperones such as ERdj4 and additional transcriptional activators such as Chop.2
The structural analysis of the luminal stress-sensing domain of yeast IRE1 provided insights into IRE1 activation under ER stress. Upon UPR activation, the luminal domain of IRE1 undergoes oligomerization, which brings the cytosolic kinase domains into juxtaposition allowing their activation by transautophosphorylation and cofactor binding. Though an otherwise conventional protein kinase, the only known phosphorylation substrate of IRE1 is itself. The trans-autophosphorylation of IRE1 somehow unmasks its effector function by triggering conformational changes that activate the RNase domain of the enzyme, which is to cleave a conserved mRNA substrate and produce the homeostatic transcription factor XBP1s. In addition, IRE1α's RNase also causes endonucleolytic decay of many ER-localized mRNAs, including those encoding chaperones, as early events culminating in apoptosis.3 IRE1 signaling attenuates after prolonged ER stress. IRE1 then enters a refractive state even if ER stress remains unmitigated. Attenuation includes dissolution of IRE1 clusters, IRE1 dephosphorylation, and decline in endoribonuclease activity. Thus IRE1 activity is governed by a timer that may be important in switching the UPR from the initially cytoprotective phase to the apoptotic mode. Moreover, it has been demonstrated that the duration of PERK and IRE1 signaling varied markedly after the imposition of protein misfolding. The sustained PERK signaling was detrimental to cell viability whereas the equivalent duration of IRE1 signaling was not, suggesting that extended PERK activity contributes to the cell death that occurs with chronic ER stress.4
The structural analysis of the luminal stress-sensing domain of yeast IRE1 provided insights into IRE1 activation under ER stress. Upon UPR activation, the luminal domain of IRE1 undergoes oligomerization, which brings the cytosolic kinase domains into juxtaposition allowing their activation by transautophosphorylation and cofactor binding. Though an otherwise conventional protein kinase, the only known phosphorylation substrate of IRE1 is itself. The trans-autophosphorylation of IRE1 somehow unmasks its effector function by triggering conformational changes that activate the RNase domain of the enzyme, which is to cleave a conserved mRNA substrate and produce the homeostatic transcription factor XBP1s. In addition, IRE1α's RNase also causes endonucleolytic decay of many ER-localized mRNAs, including those encoding chaperones, as early events culminating in apoptosis.3 IRE1 signaling attenuates after prolonged ER stress. IRE1 then enters a refractive state even if ER stress remains unmitigated. Attenuation includes dissolution of IRE1 clusters, IRE1 dephosphorylation, and decline in endoribonuclease activity. Thus IRE1 activity is governed by a timer that may be important in switching the UPR from the initially cytoprotective phase to the apoptotic mode. Moreover, it has been demonstrated that the duration of PERK and IRE1 signaling varied markedly after the imposition of protein misfolding. The sustained PERK signaling was detrimental to cell viability whereas the equivalent duration of IRE1 signaling was not, suggesting that extended PERK activity contributes to the cell death that occurs with chronic ER stress.4
REFERENCES
1. Lee, K.P.K. et al: Cell 132:89-100, 2008
2. Ron, D. & Hubbard, S.R.: Cell 132:24-6, 2008
3. Han, D. et al: Cell 138:562-75, 2009
4. Li, H. et al: proc. Natl. Acad. Sci. USA 107:16113-8, 2010
2. Ron, D. & Hubbard, S.R.: Cell 132:24-6, 2008
3. Han, D. et al: Cell 138:562-75, 2009
4. Li, H. et al: proc. Natl. Acad. Sci. USA 107:16113-8, 2010
Products are for research use only. They are not intended for human, animal, or diagnostic applications.
Параметры
Cat.No.: | CP10292 |
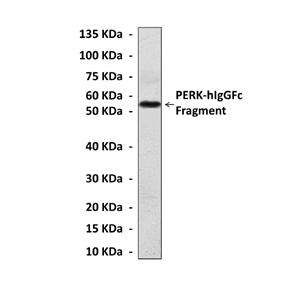
Antigen: | Raised against recombinant human PERK fragments expressed in E. coli. |
Isotype: | Mouse IgG1 |
Species & predicted species cross- reactivity ( ): | Human, Mouse, Rat |
Applications & Suggested starting dilutions:* | WB 1:1000 IP n/d IHC n/d ICC n/d FACS n/d |
Predicted Molecular Weight of protein: | 140 kDa |
Specificity/Sensitivity: | Detects endogenous PERK proteins without cross-reactivity with other family members. |
Storage: | Store at -20°C, 4°C for frequent use. Avoid repeated freeze-thaw cycles. |
*Optimal working dilutions must be determined by end user.
Документы
Информация представлена исключительно в ознакомительных целях и ни при каких условиях не является публичной офертой